Avoid the oxidative stress trap
Free radical production and oxidative stress are a mechanism of toxicity in numerous tissues and organs including liver, kidney, ear, and skin, as well as the cardiovascular and nervous systems. Mitologics’ technological platform MiToxView® offers custom-designed assays to investigate oxidative stress in whole cells and mitochondria from various sources.
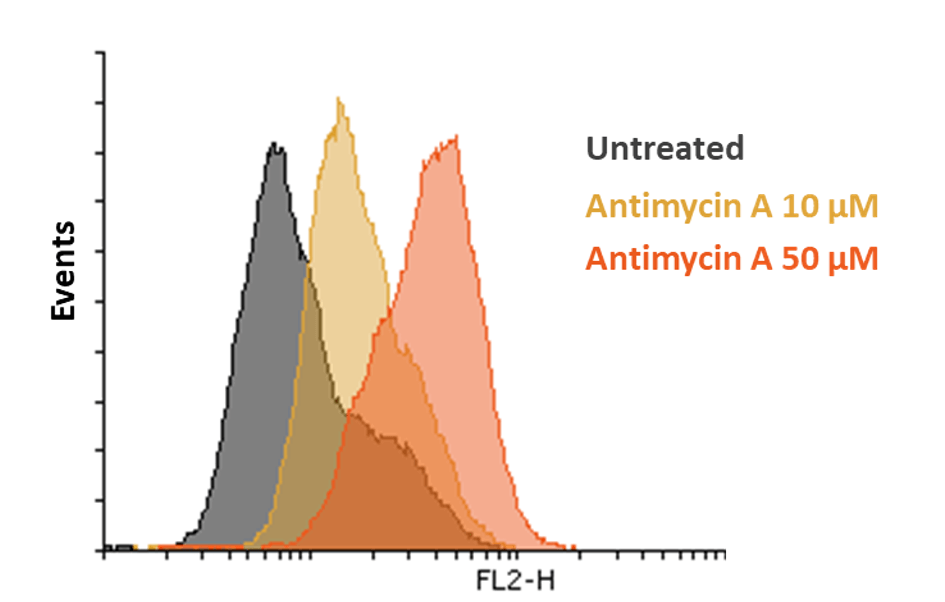
Mitochondria are major sources of reactive oxygen species (ROS) within the cell, as these molecules are produced when electrons escape through the electron carriers of the respiratory chain – complexes I, III and IV – when mitochondrial OxPHOS activity becomes altered.
As a vestigial organelle, the mitochondrion itself is very vulnerable to oxidative damage. Exposure to ROS induces lipid peroxidation, protein oxidation, and mitochondrial DNA (mtDNA) damage, creating a feedback loop that further aggravates mitochondrial dysfunction.
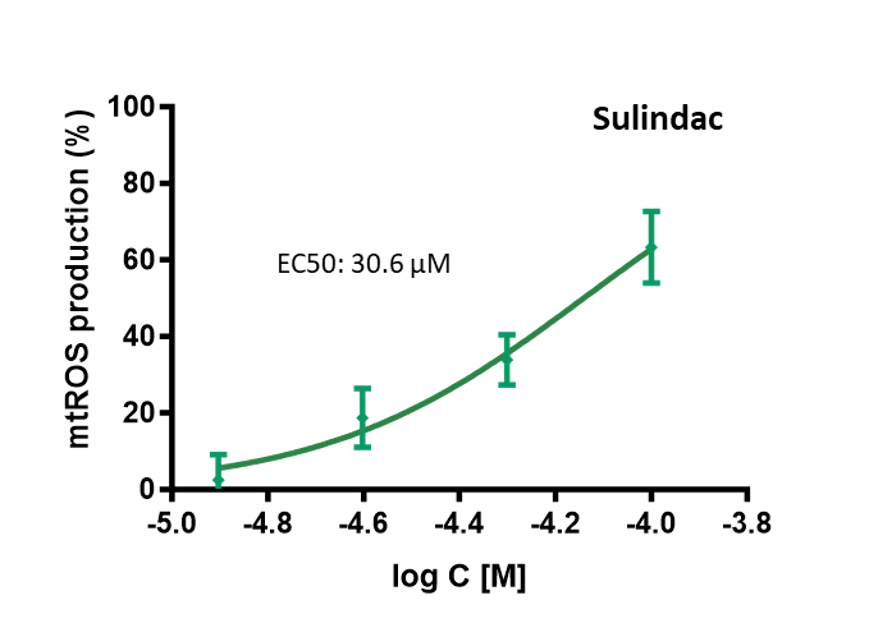
Evaluate direct effect on ROS production by mitochondria.
Induction of mtROS (superoxide anions) production on isolated mouse liver mitochondria by acute treatment with Sulindac (NSAID) showing direct mitochondrial toxicity of this hepatotoxic drug. Compound dose-response is monitored by spectrofluorimetry during 45 min in presence of MitoSox dye alowing EC50 determination.
As part of our MiToxView® platform, ROS production and the anti-oxidant defence system can be assessed in whole cells or mitochondria from various sources. The consolidated data will help you to:
- Determine potential mitochondrial toxicity of compounds,
- Screen and rank compounds based on their mitochondrial effects,
- Understand a compound’s mitochondrial toxicity mechanism and its impact at the cellular level,
- Analyse oxidative state biomarkers in pre-clinical and clinical biological samples.